The YeasTSS (Yeast Transcription Start Site) database is a primary depository of TSS data generated by Dr. Zhenguo Lin's lab at Saint Louis University. TSS data are valuable for precisely determining the 5’boundary and the 5’untranslated region (5’UTR) of protein coding genes, improving genome annotation quality, and predictions of novel genes, core promoter elements, transcription factor binding sites, and other motifs associated with transcription and inferring gene regulatory network.
The TSS data were mainly obtained by Cap analysis gene expression (CAGE), which identifies genome-wide 5’ends of transcripts at a single-nucleotide resolution and simultaneously quantifies their transcription level.
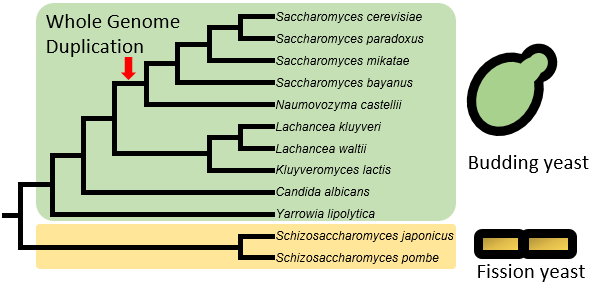
Currently, YeasTSS includes TSS data from 10 budding yeast species (Hemiascomycetes) and two fission yeast species (Schizosaccharomycetes). This database also integrates other functional genomics data related to transcription initiation and regulation, such as transcription factor binding sites (TFBS), RNA Polymerase II density and histone modification for several well-studied model yeast species, such as Saccharomyces cerevisiae, Schizosaccharomyces pombe and Candida albicans. YeasTSS enables the visualization of these functional genomics data via genome browser JBrowse. YeasTSS also provides a set of queries to search and retrieve TSS and core promoter information from gene-by-gene analysis or global approaches.
YeasTSS is updated regularly to include the latest data from my lab and recent literatures.